Latent vs. expression objective
The CD4→CD8 experiment (previous post) optimised \(\boldsymbol{\alpha}\) against the latent centroid of CD8 cells. It moves most cells in the right direction but closes gaps mainly on small-effect genes — and leaves canonical markers like CD8A/CD8B almost untouched. If we care about perturbation prediction, the thing that matters is the model’s output expression, not the layer-12 representation. Do the two agree?
Two objectives, one transition
Same contrastive setup, same CD4 source, same CD8 target — two different loss functions:
- Latent objective (previous post): minimise distance from the steered CD4 embedding to the CD8 centroid in residual-stream space.
- Expression objective (this post): minimise distance from the steered output expression to the mean CD8 output expression, weighted by log fold- change of the top DE genes.
We evaluate each run on both metrics.
Result: the two objectives are nearly orthogonal
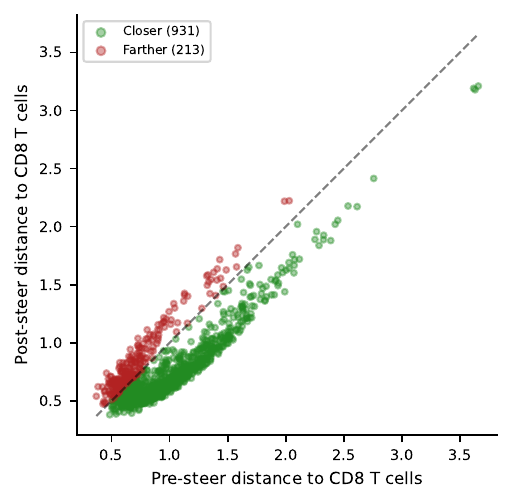
Latent objective pulls cells toward the CD8 centroid (81% closer) but barely moves expression toward CD8-like DE gene values.
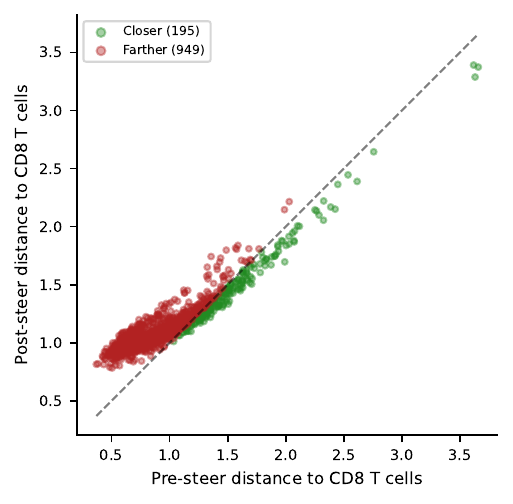
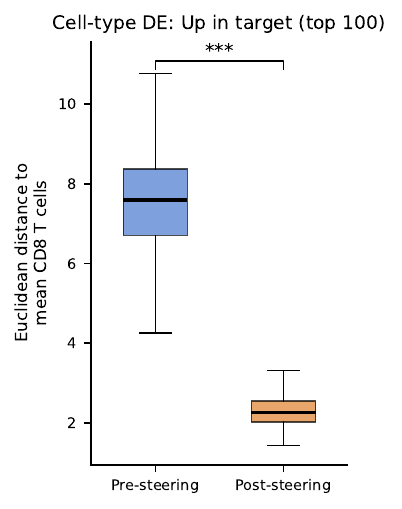
Expression objective shrinks the gap on CD8-DE genes dramatically (boxplot below) — but the same cells drift further from the CD8 centroid in latent space. Nearly the mirror image of the latent-objective scatter.
Side note: the equivalent cell-type DE boxplot for the latent objective has not been generated yet (it’s just a matter of re-running
scripts/steering/visualize_optimized_steering.pyon the latent \(\alpha\) vector — the script is objective-agnostic). I’ll drop it in once run.
What this means
The scFM’s residual-stream geometry and its output-expression geometry are not aligned. A direction in latent space that moves you to CD8’s centre does not correspond to emitting a CD8-like transcriptome — and vice versa. Two consequences:
- Care about what you optimise. If the downstream use is perturbation prediction, optimise in expression space. If the downstream use is embedding-based similarity, optimise in latent space. The two are not interchangeable.
- scFM latent similarity is not a sufficient proxy for phenotypic similarity. This is a concrete, testable caveat against the common pattern of using scFM embeddings as surrogate cell-state distances.
A follow-up observation: feature recruitment blows up
The expression objective needs ~5× more features to close the DE gap than the latent objective does (77 vs 16 features with \(|\alpha-1|>0.5\)). That hints at structure worth investigating: is closing the expression gap genuinely a higher-dimensional problem than closing the centroid gap, or is the optimiser overshooting and recruiting weak features? Diagnostic work ongoing — see reports/feature_recruitment_analysis.md.
Further reading
- Steered-feature biology under the expression objective:
reports/steered_features_biology.md. - Feature-count diagnosis:
reports/feature_recruitment_analysis.md.