Steering cell identity: CD4 → CD8
Single-feature steering works for simple programs (antiviral, BCR). But cell identity is a combination of many programs. To reprogram a CD4 T cell toward a CD8 T cell you’d expect to need more than one feature — and you want the optimizer to pick them for you, sparsely.
The contrastive objective
Let \(z(\boldsymbol{\alpha})\) be the CD4 cell’s layer-12 embedding after steering with a vector of feature scalars \(\boldsymbol{\alpha}\), and let \(\mu\) be the mean CD8 T-cell embedding (computed on the unsteered cells). Find:
\[ \min_{\boldsymbol{\alpha}} \; \lVert z(\boldsymbol{\alpha}) - \mu \rVert^2 + \lambda \lVert \boldsymbol{\alpha} \rVert_1. \]
The L1 penalty pushes \(\boldsymbol{\alpha}\) toward the no-op value (so the “steered” features are a small subset), and the squared-distance term pulls each CD4 cell toward the CD8 centroid in the SAE-reconstructed residual stream.
Result: 81% of CD4 cells move closer to CD8
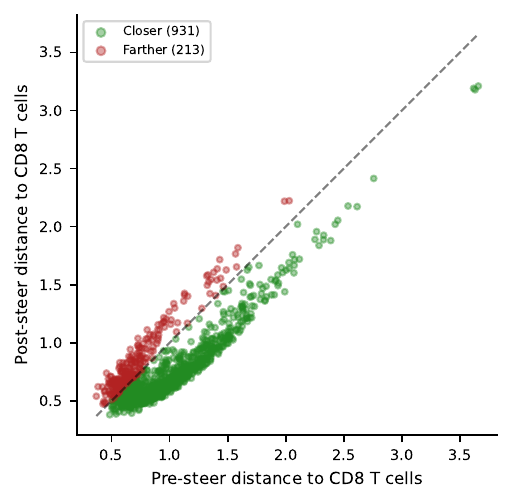
The fraction of cells that moved closer is high, but the magnitude of the moves is modest — most cells shrink their distance by a small amount, not collapse onto the CD8 centroid. Read the scatter as “the direction is right” rather than “we’ve turned CD4s into CD8s.”
On the layer-12 UMAP the CD4 cluster visibly shifts toward the CD8 cluster after steering:
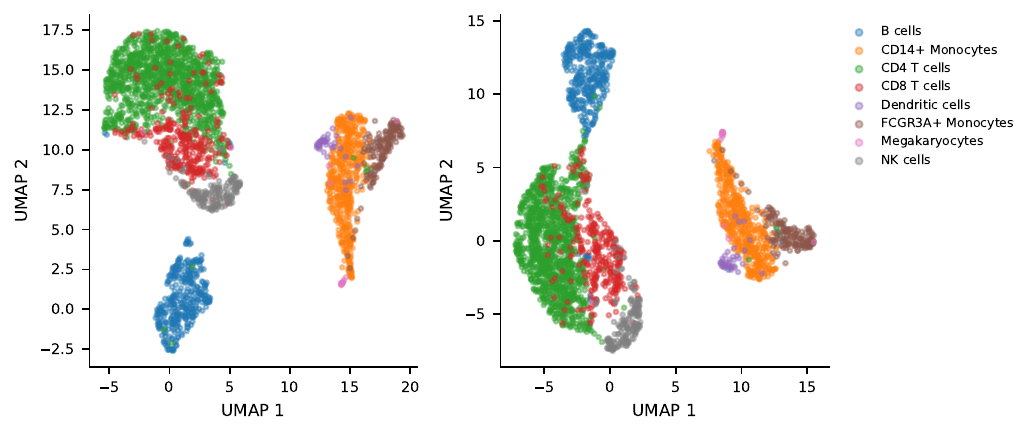
What the optimizer picked
51 features ended up with \(|\alpha - 1| > 0.3\). Union of their top-3 (by IC) GO annotations includes:
- IL-12 / IL-18 response — both promote CD8 differentiation and CTL activity.
- MHC class I antigen presentation — the class CD8s recognise (CD4s recognise MHC class II).
- CD8 α-β T cell activation and positive thymic T cell selection — direct hits.
- Cytotoxic killing of cells of other organisms and granule-mediated killing terms — CD8 effector function.
Plus a tail of generic terms (proteasome, RNA processing, housekeeping) — some noise, but the CD8-specific signal is clear and coherent. The optimiser is recruiting biologically sensible features, not arbitrary ones.
Expression-space evaluation
Latent centroid distance is one metric; output gene expression is another. Taking the intersection of CD4-vs-CD8 DE genes and post-steering DE genes, we compute each gene’s gap fraction: the share of the CD4→CD8 expression gap closed by steering.
- 70% of up-in-CD8 genes move in the correct direction.
- 65% of down-in-CD8 genes move in the correct direction.
Two caveats:
- Low-effect-size dominance. A positive effect (>50% correct direction) is only achieved once we include more than the first 100 DE genes by effect size. The steering is closing gaps predominantly on small-effect genes.
- CD8A/CD8B coreceptors remain almost intact. The canonical CD8 lineage markers are the genes you’d most hope to see flip — and they don’t. This is a pointed limitation: latent-centroid optimisation doesn’t know those specific genes “matter more” to the cell-type label.
Together these motivate the expression-space follow-up (next post).
Caveat: the objective shapes the answer
This version optimises against the layer-12 latent centroid. It works well on that metric (81% closer), but that’s the metric it was trained on. The natural follow-up is: does optimising for the latent centroid also pull the model’s output gene expression toward CD8? That’s the subject of the next post — and the answer is more surprising than I expected.
Further reading
- Full write-up with all the steered features and their GO annotations:
reports/cd4_steering.md.